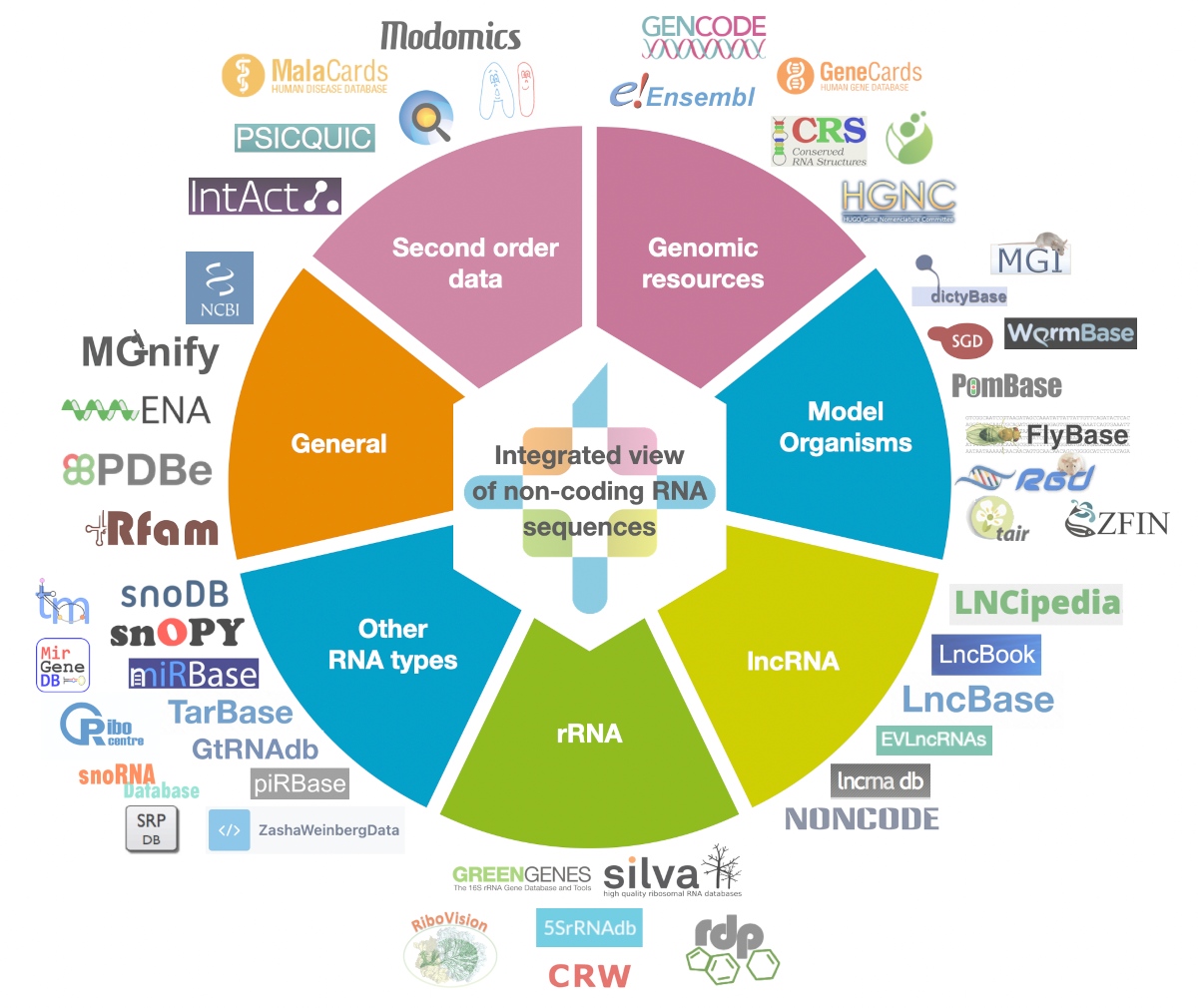
Currently the RNAcentral Consortium is formed by 61 Expert Databases, 53 of which have already been imported into RNAcentral.
If you develop an ncRNA database and would like to join RNAcentral, please contact us.
| Database | Imported | Description |
|---|---|---|
| 5SrRNAdb | 5SrRNAdb is an information resource for 5S ribosomal RNAs | |
| CRS | CRS is a database of conserved RNA motifs identified computationally in multi-species vertebrate alignments using 2D structure | |
| CRW | CRW provides comparative sequence and structure information for ribosomal, intron, and other RNAs | |
| dictyBase | dictyBase is the model organism database for the social amoeba Dictyostelium discoideum | |
| ENA | ENA is a comprehensive record of the world's nucleotide sequencing information | |
| Ensembl | Ensembl is a genome browser for vertebrate genomes and model organisms that supports research in comparative genomics, evolution, sequence variation and transcriptional regulation | |
| Ensembl Fungi | Ensembl Fungi is a genome browser for fungi genomes that complements the Ensembl database | |
| Ensembl Metazoa | Ensembl Metazoa is a genome browser for metazoan genomes that complements the Ensembl database | |
| Ensembl Plants | Ensembl Plants is a genome browser for plant genomes that complements the Ensembl database | |
| Ensembl Protists | Ensembl Protists is a genome browser for protist genomes that complements the Ensembl database | |
| Ensembl/GENCODE | GENCODE produces high quality reference gene annotation and experimental validation for human and mouse genomes | |
| EVLncRNAs | A database of lncRNAs | |
| Expression Atlas | A database of expression | |
| FlyBase | FlyBase is a database of Drosophila genes and genomes | |
| GeneCards | GeneCards is a searchable, integrative database that provides comprehensive, user-friendly information on all annotated and predicted human genes | |
| Greengenes | Greengenes is a database of full-length 16S rRNA gene that provides a curated taxonomy based on de novo tree inference | |
| GtRNAdb | GtRNAdb contains tRNA gene predictions on complete or nearly complete genomes | |
| HGNC | HGNC is the worldwide authority that assigns standardised nomenclature to human genes | |
| IntAct | IntAct provides a freely available, open source database system and analysis tools for molecular interaction data. All interactions are derived from literature curation or direct user submissions | |
| LncBase | LncBase provides experimentally verified and computationally predicted microRNA targets on long non-coding RNAs | |
| LncBook | LncBook is a curated knowledgebase of human long non-coding RNAs | |
| LNCipedia | LNCipedia is a comprehensive compendium of human long non-coding RNAs | |
| lncRNAdb | lncRNAdb is a database providing comprehensive annotations of eukaryotic long non-coding RNAs (lncRNAs) | |
| MalaCards | MalaCards integrates manually-curated and text-mining sources to associate genes, including ncRNAs, with diseases, and lists the supporting evidence | |
| MGI | MGI is the international database resource for the laboratory mouse | |
| MGnify | MGnify is a hub for the analysis and exploration of metagenomic, metatranscriptomic, amplicon and assembly data | |
| miRBase | miRBase contains high-quality miRNA annotations; miRBase is responsible for assigning official miRNA gene names | |
| MirGeneDB | MirGeneDB is a curated microRNA gene database covering 45 metazoan organisms | |
| Modomics | Modomics is a comprehensive database of RNA modifications | |
| NONCODE | NONCODE is an integrated knowledge database dedicated to non-coding RNAs | |
| PDBe | PDBe is the European repository of information about the 3D structures of large biological molecules. PDBe is a member of the Worldwide Protein Data Bank | |
| piRBase | piRBase is a database of various piRNA associated data to support piRNA functional study | |
| PLncDB | PLncDB provides comprehensive genomic view of plant lncRNAs | |
| PomBase | PomBase is a comprehensive database for the fission yeast Schizosaccharomyces pombe | |
| PSICQUIC | A database of manually annotated human miRNA interactions | |
| RDP | RDP provides quality-controlled, aligned and annotated rRNA sequences and a suite of analysis tools | |
| REDIportal | An ATLAS of A-to-I RNA editing events in human and other organisms | |
| RefSeq | RefSeq is a comprehensive, integrated, non-redundant, well-annotated set of reference sequences | |
| Rfam | Rfam is a collection of non-coding RNA families, represented by manually curated sequence alignments, consensus secondary structures and predicted homologues | |
| RGD | RGD is a rat genetic and genomic research resource | |
| Ribocentre | A database of ribozymes with representative structures | |
| RiboVision | A database of ribosomal annotations | |
| SGD | SGD provides comprehensive integrated biological information for the budding yeast | |
| SILVA | SILVA is a comprehensive resource for quality checked and aligned ribosomal RNA sequence data | |
| snoDB | snoDB is an interactive database of human snoRNA sequences, abundance and interactions | |
| snOPY | snOPY provides comprehensive information about snoRNAs, their gene loci, orthologues and their target RNAs | |
| snoRNA Database | The snoRNA Database is a curated collection of archaeal snoRNAs maintained by the Lowe Lab at UC Santa Cruz | |
| SRPDB | SRPDB provides aligned, annotated and phylogenetically ordered sequences related to structure and function of SRP | |
| TAIR | TAIR is a database of genetic and molecular biology data for the model higher plant Arabidopsis thaliana | |
| TarBase | TarBase is a collection of manually curated experimentally validated miRNA-gene interactions | |
| tmRNA Website | tmRNA Website contains predicted tmRNA sequences from RefSeq bacterial genomes, plasmids, phages and some organelles | |
| WormBase | WormBase curates, stores and displays genomic and genetic data about nematodes with primary emphasis on C. elegans and related nematodes | |
| ZFIN | The Zebrafish Information Network (ZFIN) is the database of genetic and genomic data for the zebrafish (Danio rerio) as a model organism | |
| ZWD | ZWD is a git-based collection of non-coding RNA alignments maintained by Dr Zasha Weinberg | |
| miRTarBase | miRTarBawse is an experimentally validated microRNA-target interactions database | |
| NPInter | NPInter contains data on experimentally determined functional interactions between ncRNAs and proteins, mRNAs or genomic DNA | |
| RNApathwaysDB | RNApathwaysDB contains RNA maturation and decay pathways | |
| snoRNA Atlas | snoRNA Atlas is a database of human snoRNAs | |
| sRNAmap | sRNAmap is a collection of sRNA sequences and interactions | |
| tmRDB | tmRDB is a collection of aligned, annotated and phylogenetically ordered sequences related to structure and function of tmRNA | |
| tRNAdb | tRNAdb is a compilation of tRNA sequences and tRNA genes. |